A journal is driven by the enthusiasms of its editors. Ours, fed and guided by our distinguished Editorial Board, and constrained by countless conscientious referees, are reflected in the selections that follow from the BMC Biology editorial team from the articles we published in 2016.
We don’t claim to have picked the most important ones – who could possibly judge that from the myopic perspective of twelve months? – They’re the ones each of the team found most interesting, for any of many reasons, or was proudest of having published – or indeed recognized as being important to one of the great issues in biology, from human disease and the defeat of its vectors to crop security in a warming world.
There are four attributed contributions, one from each of the four editors, identified by the picture at the start of each selection.
But before I hand over to the team, I have two articles to add to the selection.
One is James Edgar’s Q&A on exosomes. These baffling extracellular vesicles have been credited with roles in the transmission of immunity and disease, and (conversely) with potential for drug delivery. But their biology remains largely a cipher. If you have been looking for somewhere to get your bearings on this vexed issue, try James’s piece.
My other choice is Eugene Koonin’s assault on the ‘Just-so’ approach to evolutionary biology – this is not just a re-run of Gould and Lewontin and the famous spandrels: it’s Koonin’s perspective on how non-adaptive features come about – or, as he has it, ‘the splendor and the misery of…the neutral null’.
Both articles, in different ways, reflect, court, address controversy. We embrace controversy – not for its own sake, but for the energy it brings to the development of ideas and ultimately to advances in understanding.
Enough. Over to the team.
 Thomas Wylie – Assistant Editor
Thomas Wylie – Assistant Editor
If you want to understand function…
Francis Crick is alleged to have said (and did not deny having said), “If you want to understand function, study structure.” Many biological issues were tackled by structural biology papers we published this year. From informing strategies to counter of the deepening problem of resistance to the last ditch antibiotic colistin via the structure of the resistance enzyme to studies using structure to study the mechanisms of prophenoloxidase activation in the mosquito immune system, substrate recognition by the bacterial dual chaperone/PPI SlyD and the molecular basis for a form of CMT disease, and resistance of acute lymphoblastic leukamia to chemotherapy.
“If you want to understand function, study structure.”
The Daumke group’s paper in February correcting their previous hypothesis on Irga6 dimerization on the basis of new evidence from the stucture of the dimer was a personal highlight, as it was the first paper I was assigned in my role at BMC Biology, as well as, I thought, a nice example of how science corrects itself.
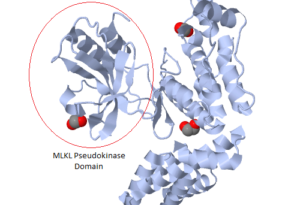
Another highlight for me was the conferences I reported on: the inaugural Astbury Conversation in April and a meeting on Pseudoenzymes in September were great for different reasons, the first showing the breadth of work and progress in structural biology, and the other a fascinating look at how these catalytically deficient enzymes have evolved and provide insights into new ways to target signaling pathways for therapeutic purposes.

Graham Bell – Senior Editor
Breaking the law? Evolution in reverse
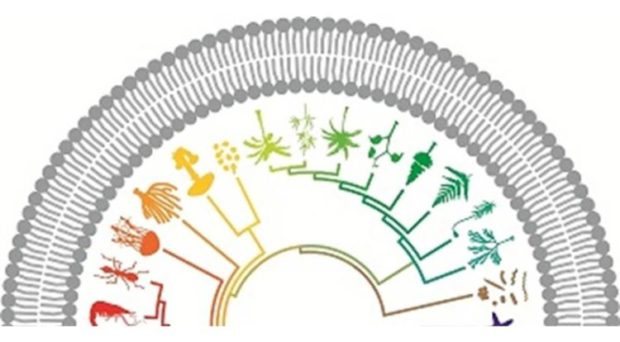
One of the papers I enjoyed reading most was from Jan Andersson and colleagues on the reversibility of parasitism and Dollo’s ‘law’, which suggests that evolution is not reversible – complex traits, once lost, cannot re-evolve the same way. As organisms change from a free-living to a parasitic lifestyle over evolutionary time, they undergo adaptations to living with, on, or in, hosts.
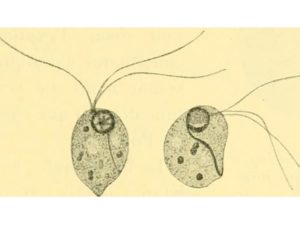
These adaptations to a parasitic lifestyle – losses of features like digestive enzymes for feeding, and gains of other functions – are generally thought to be irreversible – or so the textbooks say. But the paper suggests that this isn’t true: biology is more ingenious, and the genes needed to restore these these functions can be reintroduced from other species by horizontal gene transfer.
Adaptations to a parasitic lifestyle are generally thought to be irreversible – or so the textbooks say
We’ve published a few HGT papers this year – and received submissions of many more – on what is clearly still a highly contentious topic in need of cool and level-headed analysis. Chuan Ku and Bill Martin offered their “70% rule” as a guide for when claims of HGT might be reasonable, with an associated commentary from Etienne Danchin to provide perspective.
What is wrong with this picture?
To give a bit of a cheat answer on articles to highlight, working on and hearing the feedback on our “What is wrong with this picture?” series has been particularly rewarding. The series explores some of the ways figures or statistics can be misleading – including what p values really tell us, complications from 3D graphs, and problems quantifying qualitative data like Westerns.
Our Guest Editors for the series are superb, providing sage advice and ideas, with John Brookfield contributing one of the pieces in the collection; and the feedback from the community – both emailed comments and from talking to people at meetings – has been positive and constructive. We’ll be continuing the series in 2017 and are open to suggestions for future articles.
Life in the dark
One of our first papers published in 2016 was the genomes of 3 cavefish species, adapted to living in different levels of light: from surface dwellers to those who inhabit pitch black caves. As an emerging model organism this looks like becoming an important resource, and the level of analysis seemed really impressive. (It also gave rise to one of my favourite reviews, where a ferociously conscientious referee enthused about some “very interesting’” results that the authors perhaps hadn’t quite appreciated, hidden in the depths of what were originally Figures S28 and S30). We hope to have much more on model (and ‘non-model’!) organisms this year.
 Christian Matheou – Associate Editor
Christian Matheou – Associate Editor
Securing our crops
More genomes and genomics came from the plant world and its pests and pathogens. Book-marking the other end of the year, we published a draft genome for the whitefly Bemisia tabaci, a major disease vector for crops. The B tabaci genome provides insight into the success of the whitefly as an invasive agricultural pest (innovations such as the expansion of gene families involved in detoxification, and horizontal gene transfer from fungi and bacteria), which could help suggest better ways to control it.
Whitefly carry plant pathogenic viruses – but with climate change, the increasing prevalence of fungal diseases is a growing concern. February 2016 saw the emergence of a devastating Bangladeshi fungus that has since led to up to 90% wheat yield losses, from over 15,000 hectares. A major collaborative effort (31 researchers from 14 institutes) identified the fungus as a wheat-infecting lineage of the Magnaporthe oryzae fungus, most probably originating in South America, on the basis of a phylogenomic and population genomic analyses (the raw data from which were released as soon as they were available). This suggests that methods– such as disease resistant cultivars and fungicides – that are used to combat M oryzae in Brazil might be directly applied to the Bangladeshi outbreak.
With climate change, the increasing prevalence of fungal diseases is a growing concern
Disease resistance is for many obvious reasons preferable to chemical control, and the innate immune devices of plants are a promising resource for new sources of resistance to be exploited for crops. One trick plants have for identifying pathogens is to fuse their immune receptors to domains resembling pathogen targets. This manoevre is described by Jeffrey Ellis in a Commentary on a paper published at the beginning of last year as part of our Plant Signals series. In an extensive search across the genomes of 40 plants, Ksenia Krasileva and colleagues identified 265 different domains fused to plant immune receptors. This provides a valuable resource for identifying potential targets of plant pathogens, and possibly new sources of disease resistance.
Plant signals
Our Plant Signals series has grown a lot this year under our guest editors Elliot Meyerowitz, Virginia Walbot and Wolf Frommer. The aim of the series is to provide an introduction to the key signalling molecules and pathways of plants and their functions for non-specialists – highlighting what we know, as well as what we (currently) don’t.
Just over a year ago, we published a Q&A from Steven Smith and colleagues on karrikins – signalling molecules that are produced from burning plant material, and that can stimulate germination in a variety of plants. Most notable are the ‘fire-followers’ that remain dormant until a wild-fire releases the required karrikins and triggers germination.
Contributions since then include Caren Chang on ethylene, and its current and potential future application in agriculture; Maike Breiden and Rudiger Simon on peptide signalling, and its role in plant development; Sebastien Paque and Dolf Weijers on Auxin, and how it does (almost) everything, Antoine Larrieu and Teva Vernoux on jasmonate, and it’s importance for plant adaptation; and Jijie Chai and colleagues discussing brassinosteroids, which, like the animal steroid hormones they resemble, regulate a diverse range of plant biology.
We’re looking forward to to extending the series in 2017, and will also be welcoming relevant research submissions.
 Penelope Austin – Deputy Editor
Penelope Austin – Deputy Editor
A forum on memory and a convocation of worms
“The mechanism of memory remains one of the great unsolved problems of biology”. So started our forum article that gathered together contemporary perspectives on this question from leading researchers in the field. Seeing this published in May 2016, and its enthusiastic reception by readers, was a major highlight of my year, so thanks to Guest Editor and Editorial Board member Mu Ming Poo for support and guidance, and – not least – his own contribution.
“The mechanism of memory remains one of the great unsolved problems of biology”
Another memorable 2016 article describes the native microbiome of the nematode Caenorhabditis elegans, setting the stage for studies on the impact of a natural microbiome on the biology of this highly successful model organism, an achievement put in perspective in an accompanying commentary from Laura Clarke and Jonathan Hodgkin. I don’t know what these roundworms did to deserve the label elegans , but assessing papers for BMC Biology has taught me to appreciate these simple creatures for their contribution to science, and the field for its co-operative collegiality.
The worm microbiome paper was just one of a convocation of worms on our website in 2016, including a paper from Bob Horvitz on a new mechanism of cell death – “assisted suicide” – that requires engulfment by phagocytosis as well as the expression of apoptotic pathway genes. Of the others, most illustrated the remarkable ability of this simple invertebrate to yield insights into the function of conserved genes relevant to human disease. For example, a paper from Nathalie Pujol and colleagues identified an essential role for the pseudokinase tribbles, whose misfunction is associated with leukaemia, metabolic syndromes and cardiovascular disease, in suppressing the conserved transcription factor C/EBP and preventing the activation of innate immune pathways during C. elegans development, (while a companion paper from Fred Ausubel and colleagues reported evidence for its post-developmental role in intestinal immune surveillance.)
A paper that kindled my sympathy for worms was from Andy Biron and colleagues on the role of serotonin in accelerating their decision making. Yes C. elegans needs serotonin too, and this study demonstrated a clear quantitative effect of serotonin deficiency – a slowed deceleration on encountering a patch of food – that impaired their ability to forage effectively when food was scarce.
But it’s not just as a model organism that nematodes make their appearance in our pages. A paper from Elissa Hallem and colleagues, on the temperature-seeking behaviour of parasitic nematodes, reflects our interest in their wider ecological relevance and use as biocontrol agents against insect pests – warfare on pests and pathogens being a recurrent theme in BMC Biology: see contributions to this blog above!
Miranda Robertson
Latest posts by Miranda Robertson (see all)
- The scientific Odyssey: Pre-registering the voyage - 3rd May 2017
- A year of almost anything you can think of in life science – an eclectic pick from BMC Biology - 13th January 2017
- Peer review: opting out - 23rd September 2016
Comments